PhD Top Stories
Anna Schroeder
DNA metabarcoding and morphological analysis - Assessment of zooplankton biodiversity in transitional waters
Doctoral Programme in Environmental Life Sciences
Zooplankton biodiversity assessment is a crucial element in monitoring marine ecosystem processes and community responses to environmental alterations. As accurate morphological assessments are labour-intensive, the estimation of biodiversity with DNA metabarcoding using high-throughput sequencing (HTS) is becoming an important tool for surveying biodiversity (Taberlet et al., 2012). In fact, this technique has the advantage of a broad taxonomic coverage paired with an elevated sample processing speed, allowing to increase the sampling effort (frequency and spatial coverage) with sustainable costs (Brannock et al., 2014; Coissac et al., 2012). This study aims to evaluate the suitability of metabarcoding for zooplankton biodiversity assessment and biomonitoring in a transitional environment. Therefore, seasonal zooplankton sampling was carried out in the Venice Lagoon and the nearby coastal area (Northern Adriatic Sea) and samples were divided in two equal parts: one for taxonomic and quantitative determinations performed by stereomicroscope, and the other part for genetic zooplankton community analysis utilising a fragment (313 bp) of the cytochrome c oxidase subunit I (COI) (Leray et al., 2013).
The molecular analysis showed to be effective in reflecting the complexity of zooplankton assemblages, as the taxa richness resulted to be higher compared to the classical morphological method (224 vs. 88 taxa). These differences highlight the power of this method in discriminating cryptic and sibling species and the meroplanktonic component, morphologically identified only up to order level. Nevertheless these differences, the ecological evaluation of both methods showed comparable outcomes. The analysis of alpha diversity, measured by the Shannon-Wiener Index, gave similar and significantly correlated results for metabarcoding and morphological identification. Moreover, the two methods show a similar spatio-temporal pattern, presenting a separation by seasons, following a gradient in temperature, and by location following the sea-lagoon salinity gradient (sea, inlet, lagoon) typical for transitional waters.(Fig.1)

Figure 1: Beta diversity estimates based on Bray-Curtis similarities of molecular (MBC) and morphological (MOI) data, respectively. Colours of points refer to the sampling season of each sample, while the three locations (sea, inlet, lagoon) are highlighted plotting the distance to their centroid and the standard deviations of the points per location with the respective colours. Salinity and temperature are superimposed according to the CTD measurements during sampling.
However, for both biodiversity measures it has to be taken into account that the metabarcoding data is based on the number of reads as a proxy of biomass (Lindeque et al., 2013), while the morphological identification was based on individual counts. Correlations between sequence data and species abundance have been a focus of several studies, but in general low associations between abundance or biomass and read number have been obtained (e.g. Harvey et al., 2017). But, similarly to the findings of Bucklin et al. (2019) for the 18S (V9) marker, in this study, the COI marker was shown to be very promising when it comes to the quantification of important taxonomic groups and a variety of taxa. The numbers of sequences and abundance counts based on morphological taxonomic identifications were significantly correlated for selected species (present in both datasets), for the most abundant classes of arthropods and for most phyla (Fig. 2).
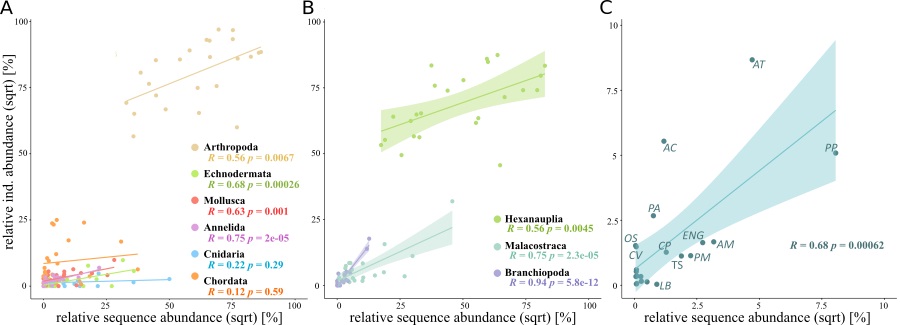
Figure 2: Relative abundance of reads for molecular data and of individual counts or morphological data of (A) most abundant phyla; (B) most abundant classes of Arthropoda and (C) 21 selected species present in both datasets (AT: Acartia tonsa, AC: Acartia clausi, PP: Paracalanus parvus, PA: Penilia avirostris, Eng: Engraulis sp., OS: Oithona similis, CV: Ctenocalanus vanus, CP: Centropages ponticus, TS: Temora stylifera, PM: Pseudodiaptomus marinus, LB: Labidocera brunescens; species with lowest abundances are not labelled. Pearson correlations between the two methods are given in the corresponding colour).
Our results indicate that DNA metabarcoding is an efficient tool for biodiversity assessments especially in ecosystems with high spatial and temporal variability, where high sampling effort is required and where high spatio-temporal coverage is preferred over the information on population structure and it is particularly useful e.g. when studying larval dispersion patterns as well as a fast alert systems for non-native species (NIS).
Authors and affiliations
Anna Schroeder1,2, David Stankovic3, Alberto Pallavicini2,4, Fabrizia Gionechetti2, Marco Pansera1, Elisa Camatti12University of Trieste, Department of Life Sciences, Trieste, Italy
3Marine Biology Station Piran, National Institute of Biology, Piran, Slovenia
4Stazione zoologica Anton Dohrn, Naples, Italy
Contact
Anna Schroeder, email: anna.schroeder@phd.units.itReference
A. Schroeder, D. Stankovic, A. Pallavicini, F. Gionechetti, M. Pansera and E. CamattiDNA metabarcoding and morphological analysis-Assessment of zooplankton biodiversity in transitional waters.
Marine Environmental Research 160, 104946 (2020)
https://www.sciencedirect.com/science/article/pii/S0141113619308852
Informazioni aggiornate al: 03.3.2021 alle ore 12:26
Contact: Webmaster - Università di Trieste pagina curata da: Research Doctorate

Piazzale Europa, 1 - 34127 - Trieste, Italia -
Tel. +39 040 558 7111 - P.IVA 00211830328
C.F. 80013890324 - P.E.C. ateneo@pec.units.it



